Prioritization of genes driving congenital phenotypes of patients with de novo genomic structural variants.
Authors of this article are:
Middelkamp S, Vlaar JM, Giltay J, Korzelius J, Besselink N, Boymans S, Janssen R, de la Fonteijne L, van Binsbergen E, van Roosmalen MJ, Hochstenbach R, Giachino D, Talkowski ME, Kloosterman WP, Cuppen E.
A summary of the article is shown below:
BACKGROUND: Genomic structural variants (SVs) can affect many genes and regulatory elements. Therefore, the molecular mechanisms driving the phenotypes of patients carrying de novo SVs are frequently unknown.METHODS: We applied a combination of systematic experimental and bioinformatic methods to improve the molecular diagnosis of 39 patients with multiple congenital abnormalities and/or intellectual disability harboring apparent de novo SVs, most with an inconclusive diagnosis after regular genetic testing.RESULTS: In 7 of these cases (18%), whole-genome sequencing analysis revealed disease-relevant complexities of the SVs missed in routine microarray-based analyses. We developed a computational tool to predict the effects on genes directly affected by SVs and on genes indirectly affected likely due to the changes in chromatin organization and impact on regulatory mechanisms. By combining these functional predictions with extensive phenotype information, candidate driver genes were identified in 16/39 (41%) patients. In 8 cases, evidence was found for the involvement of multiple candidate drivers contributing to different parts of the phenotypes. Subsequently, we applied this computational method to two cohorts containing a total of 379 patients with previously detected and classified de novo SVs and identified candidate driver genes in 189 cases (50%), including 40 cases whose SVs were previously not classified as pathogenic. Pathogenic position effects were predicted in 28% of all studied cases with balanced SVs and in 11% of the cases with copy number variants.CONCLUSIONS: These results demonstrate an integrated computational and experimental approach to predict driver genes based on analyses of WGS data with phenotype association and chromatin organization datasets. These analyses nominate new pathogenic loci and have strong potential to improve the molecular diagnosis of patients with de novo SVs.
Check out the article’s website on Pubmed for more information:
[link-preview url=https://www.ncbi.nlm.nih.gov/pubmed/31801603 forceshot=true]
This article is a good source of information and a good way to become familiar with topics such as: Copy number variants; Driver genes; Intellectual disability; Multiple congenital anomalies; Neurodevelopmental disorders; Position effects; Structural variation; Topologically associating domains; Transcriptome sequencing; Whole-genome sequencing.
NativeFolder: The Only Bacterial Culture Medium for the Expression of Soluble Proteins
-
- Sale!
- Molecular Biology, Research Kits
NativeFolder Bacterial Culture Medium
- Original price was: $795.00.$295.00Current price is: $295.00.
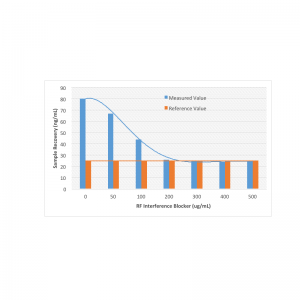
Interference Test Kit for Assay Validation
-
- Sale!
- Interference, Research Kits
Interference Test Kit
- Original price was: $995.00.$495.00Current price is: $495.00.
Rheumatoid Factor Interference Blocker
-
- Sale!
- Antibodies, Interference
Rheumatoid Factor Interference Blocker
- Original price was: $595.00.$395.00Current price is: $395.00.
New Antibodies from MOLECULAR DEPOT
-
- Sale!
- Antibodies, Biochemistry
Rabbit Polyclonal CTNL1 CTNNAL1 Antibody (Human, Mouse)
- Original price was: $1,595.00.$795.00Current price is: $795.00.
-
- Sale!
- Antibodies, Biochemistry
Rabbit Monoclonal CXCL11 Antibody
- Original price was: $1,595.00.$795.00Current price is: $795.00.
-
- Sale!
- Antibodies, Biochemistry
Rabbit Polyclonal Cullin 4a (CUL4A) Antibody (Human)
- Original price was: $1,595.00.$795.00Current price is: $795.00.
-
- Sale!
- Antibodies, Biochemistry
Rabbit Polyclonal Cullin 1/CUL1 (N-Terminus) Antibody (Human, Mouse, Rat)
- Original price was: $1,595.00.$795.00Current price is: $795.00.
-
- Sale!
- Antibodies, Biochemistry
Rabbit Polyclonal CTNA2 CTNNA2 Antibody (Human, Mouse)
- Original price was: $1,595.00.$795.00Current price is: $795.00.
-
- Sale!
- Antibodies, Biochemistry
Rabbit Monoclonal CTR1/SLC31A1 Antibody
- Original price was: $1,595.00.$795.00Current price is: $795.00.
-
- Sale!
- Antibodies, Biochemistry
Mouse Monoclonal CTLA4 Antibody
- Original price was: $1,795.00.$895.00Current price is: $895.00.
-
- Sale!
- Antibodies, Biochemistry
Rabbit Polyclonal CXCL5 (A38-Q130) Antibody (Rat)
- Original price was: $1,595.00.$795.00Current price is: $795.00.
-
- Sale!
- Antibodies, Biochemistry
Rabbit Polyclonal CXCR1 (N-Terminus) Antibody (Human)
- Original price was: $1,595.00.$795.00Current price is: $795.00.
-
- Sale!
- Antibodies, Biochemistry
Anti-CXB5 GJB5 Antibody
- Original price was: $1,595.00.$795.00Current price is: $795.00.
New Proteins from MOLECULAR DEPOT
-
- Sale!
- Conjugates, Proteins
Captopril HSA Biotin Conjugate
- Original price was: $1,995.00.$995.00Current price is: $995.00.
-
- Sale!
- Conjugates, Proteins
Aminopenicillic Acid HRP Conjugate
- Original price was: $1,595.00.$795.00Current price is: $795.00.
-
- Sale!
- Biochemistry, Proteins
PKC-α, Active, GST-tagged from Xanopus sp.
- Original price was: $1,595.00.$795.00Current price is: $795.00.
-
- Sale!
- Conjugates, Proteins
Prilocain HSA Biotin Conjugate
- Original price was: $1,995.00.$995.00Current price is: $995.00.
-
- Sale!
- Conjugates, Proteins
T3 (Triiodothyronine) HSA Biotin Conjugate
- Original price was: $1,995.00.$995.00Current price is: $995.00.
-
- Sale!
- Conjugates, Proteins
Pipemidic acid HSA Biotin Conjugate
- Original price was: $1,995.00.$995.00Current price is: $995.00.
-
- Sale!
- Biochemistry, Proteins
PKD2 Protein, Active (Recombinant Human)
- Original price was: $1,795.00.$895.00Current price is: $895.00.
-
- Sale!
- Conjugates, Proteins
Meropenem BSA Biotin Conjugate
- Original price was: $1,995.00.$995.00Current price is: $995.00.
-
- Sale!
- Conjugates, Proteins
Piperacillin HSA Biotin Conjugate
- Original price was: $1,995.00.$995.00Current price is: $995.00.
-
- Sale!
- Biochemistry, Proteins
Protein A Peroxidase from Staphylococcus aureus/horseradish
- Original price was: $1,195.00.$595.00Current price is: $595.00.
New Chemicals from MOLECULAR DEPOT
-
- Sale!
- Chemicals, Interference, Lipids
Triglyceride Mix for Interference Testing
- Original price was: $895.00.$395.00Current price is: $395.00.
-
- Sale!
- Chemicals
Microparticle Stabilizer Solution
- Price range: $295.00 through $495.00
-
- Sale!
- Buffers & Solutions, Chemicals
Enzyme Acceptor Stabilization Buffer
- Original price was: $495.00.$250.00Current price is: $250.00.
-
- Sale!
- Buffers & Solutions, Chemicals
Enzyme Donor Stabilization Buffer
- Original price was: $495.00.$250.00Current price is: $250.00.
-
- Sale!
- Chemicals, Conjugates
Progesterone Biotin Conjugate Solution
- Original price was: $650.00.$350.00Current price is: $350.00.
-
- Sale!
- Buffers & Solutions, Chemicals
Microparticle Activation Buffer
- Price range: $295.00 through $495.00
-
- Sale!
- Buffers & Solutions, Chemicals
Microparticle Washing Buffer
- Price range: $295.00 through $495.00
-
- Sale!
- Buffers & Solutions, Chemicals
Microparticle Blocking Buffer
- Price range: $395.00 through $695.00
-
- Sale!
- Buffers & Solutions, Chemicals
Latex Microparticles Anti Aggregation Agent
- Price range: $295.00 through $3,500.00
-
- Sale!
- Chemicals
CPRG (Chlorophenol red-β-D-galactopyranoside)
- Original price was: $850.00.$395.00Current price is: $395.00.